 y.algo.Groups
y.algo.Groups
|
Search this API | ||||||||
| PREV CLASS NEXT CLASS | FRAMES NO FRAMES | ||||||||
| SUMMARY: NESTED | FIELD | CONSTR | METHOD | DETAIL: FIELD | CONSTR | METHOD | ||||||||
java.lang.Objecty.algo.Groups
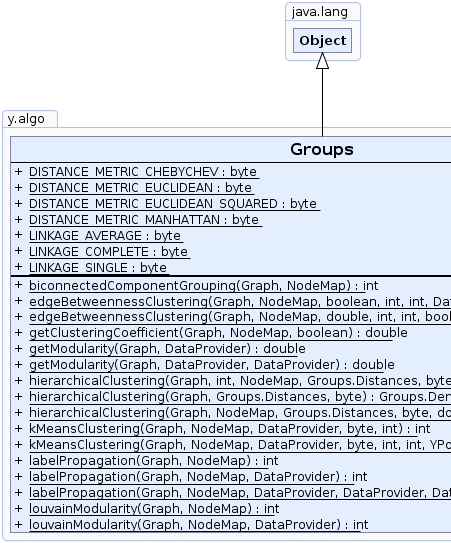
public class Groups
This class provides methods for automatically partitioning nodes of a graph into groups.
Partitions can be defined using edge betweenness centrality, biconnectivity, k-means clustering, hierarchical clustering or community detection algorithms like label propagation or louvain modularity.
This class also contains methods that provide clustering-related information like the average clustering coefficient, the average weighted coefficient and the modularity of a graph network.
k-clusters based on their
positions on the plane and a given distance metric.
Our implementation is based on the description in: "Near linear time algorithm to detect community structures in large-scale networks" by U. N. Raghavan, R. Albert, and S. Kumara in Phys. Rev. E 76, 036106, 2007
Our implementation is based on the description in: "Fast unfolding of communities in large networks" by V.D. Blondel, J.L. Guillaume, R. Lambiotte and E. Lefebvre, in Journal of Statistical Mechanics: Theory and Experiment 2008 (10), P10008 (12pp).
1 indicates that the node's neighbors form a clique (i.e., there is an edge between each
pair of neighbors).
 |
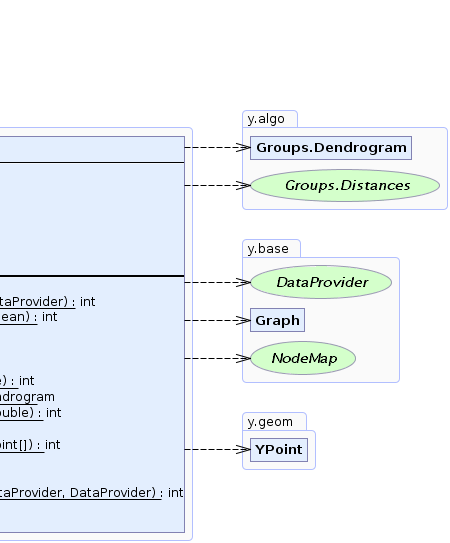 |
| Nested Class Summary | |
|---|---|
static class |
Groups.Dendrogram
This class provides the result of hierarchical clustering algorithms by means of a binary tree structure. |
static interface |
Groups.Distances
An interface that determines the distance between two nodes of a graph. |
| Field Summary | |
|---|---|
static byte |
DISTANCE_METRIC_CHEBYCHEV
A specifier for Chebychev distance metric. |
static byte |
DISTANCE_METRIC_EUCLIDEAN
A specifier for euclidean distance metric. |
static byte |
DISTANCE_METRIC_EUCLIDEAN_SQUARED
A specifier for euclidean squared distance metric. |
static byte |
DISTANCE_METRIC_MANHATTAN
A specifier for Manhattan distance metric. |
static byte |
LINKAGE_AVERAGE
A specifier for average-linkage (or mean-linkage) clustering. |
static byte |
LINKAGE_COMPLETE
A specifier for complete-linkage (or maximum-linkage) clustering. |
static byte |
LINKAGE_SINGLE
A specifier for single-linkage (or minimum-linkage) clustering. |
| Method Summary | |
|---|---|
static int |
biconnectedComponentGrouping(Graph graph,
NodeMap groupIDs)
This method partitions the graph by analyzing its biconnected components. |
static int |
edgeBetweennessClustering(Graph graph,
NodeMap clusterIDs,
boolean directed,
int minGroupCount,
int maxGroupCount,
DataProvider edgeCosts)
Partitions the graph into groups using edge betweenness centrality. |
static int |
edgeBetweennessClustering(Graph graph,
NodeMap clusterIDs,
double qualityTimeRatio,
int minGroupCount,
int maxGroupCount,
boolean refine)
Partitions the graph into groups using edge betweenness clustering proposed by Girvan and Newman. |
static double |
getClusteringCoefficient(Graph graph,
NodeMap coefficientMap,
boolean directed)
Calculates the local clustering coefficient for each node and returns the average clustering coefficient. |
static double |
getModularity(Graph graph,
DataProvider communityIndex)
Computes the modularity of a given graph. |
static double |
getModularity(Graph graph,
DataProvider communityIndex,
DataProvider edgeWeight)
Computes the modularity of a given graph. |
static Groups.Dendrogram |
hierarchicalClustering(Graph graph,
Groups.Distances distances,
byte linkage)
Partitions the graph into clusters based on hierarchical clustering. |
static int |
hierarchicalClustering(Graph graph,
int maxCluster,
NodeMap clusterIDs,
Groups.Distances distances,
byte linkage)
Partitions the graph into clusters based on hierarchical clustering, while the dendrogram is cut based on a given maximum number of clusters. |
static int |
hierarchicalClustering(Graph graph,
NodeMap clusterIDs,
Groups.Distances distances,
byte linkage,
double cutOff)
Partitions the graph into clusters based on hierarchical clustering, while the dendrogram is cut based on a given cut-off value. |
static int |
kMeansClustering(Graph graph,
NodeMap clusterIDs,
DataProvider nodePositions,
byte distanceMetric,
int k)
Like kMeansClustering(Graph, NodeMap, DataProvider, byte, int, int, YPoint[]), but no initial centroids are
given as input. |
static int |
kMeansClustering(Graph graph,
NodeMap clusterIDs,
DataProvider nodePositions,
byte distanceMetric,
int k,
int iterations,
YPoint[] centroids)
Partitions the graph into clusters using k-means clustering algorithm. |
static int |
labelPropagation(Graph graph,
NodeMap finalLabel)
Detects the communities in the specified input graph by applying a label propagation algorithm. |
static int |
labelPropagation(Graph graph,
NodeMap finalLabel,
DataProvider initialLabel)
Detects the communities in the specified input graph by applying a label propagation algorithm. |
static int |
labelPropagation(Graph graph,
NodeMap finalLabel,
DataProvider initialLabel,
DataProvider nodeWeight,
DataProvider edgeWeight,
DataProvider edgeDirectedness)
Detects the communities in the specified input graph by applying a label propagation algorithm. |
static int |
louvainModularity(Graph graph,
NodeMap communityIndex)
Detects the communities in the specified input graph by applying the louvain modularity method. |
static int |
louvainModularity(Graph graph,
NodeMap communityIndex,
DataProvider edgeWeight)
Detects the communities in the specified input graph by applying the louvain modularity method. |
| Methods inherited from class java.lang.Object |
|---|
clone, equals, finalize, getClass, hashCode, notify, notifyAll, toString, wait, wait, wait |
| Field Detail |
|---|
public static final byte DISTANCE_METRIC_EUCLIDEAN
kMeansClustering(Graph, NodeMap, DataProvider, byte, int),
kMeansClustering(Graph, NodeMap, DataProvider, byte, int, int, YPoint[]),
Constant Field Valuespublic static final byte DISTANCE_METRIC_EUCLIDEAN_SQUARED
kMeansClustering(Graph, NodeMap, DataProvider, byte, int),
kMeansClustering(Graph, NodeMap, DataProvider, byte, int, int, YPoint[]),
Constant Field Valuespublic static final byte DISTANCE_METRIC_MANHATTAN
kMeansClustering(Graph, NodeMap, DataProvider, byte, int),
kMeansClustering(Graph, NodeMap, DataProvider, byte, int, int, YPoint[]),
Constant Field Valuespublic static final byte DISTANCE_METRIC_CHEBYCHEV
kMeansClustering(Graph, NodeMap, DataProvider, byte, int),
kMeansClustering(Graph, NodeMap, DataProvider, byte, int, int, YPoint[]),
Constant Field Valuespublic static final byte LINKAGE_SINGLE
The distance between two clusters is determined by the minimum distance. Clusters thus are as close as their closest pair of nodes.
hierarchicalClustering(Graph, Distances, byte),
hierarchicalClustering(Graph, int, NodeMap, Distances, byte),
hierarchicalClustering(Graph, NodeMap, Distances, byte, double),
Constant Field Valuespublic static final byte LINKAGE_COMPLETE
The distance between two clusters is determined by the maximum distance. Clusters thus are as close as the pair of nodes that are furthest away from each other.
hierarchicalClustering(Graph, Distances, byte),
hierarchicalClustering(Graph, int, NodeMap, Distances, byte),
hierarchicalClustering(Graph, NodeMap, Distances, byte, double),
Constant Field Valuespublic static final byte LINKAGE_AVERAGE
The distance between two clusters is the average distance between each pair of nodes.
hierarchicalClustering(Graph, Distances, byte),
hierarchicalClustering(Graph, int, NodeMap, Distances, byte),
hierarchicalClustering(Graph, NodeMap, Distances, byte, double),
Constant Field Values| Method Detail |
|---|
public static int edgeBetweennessClustering(Graph graph,
NodeMap clusterIDs,
boolean directed,
int minGroupCount,
int maxGroupCount,
DataProvider edgeCosts)
In each iteration the edge with the highest betweenness centrality is removed from the graph. The method stops, if there are no more edges to remove. The clustering with the best quality reached during the process will be returned.
The method requires the maximum number of groups that will be returned. The smaller this value is, the
faster the overall computation time.
The upper bound on the number of groups is graph.N(). Also, the number of returned groups is
never smaller than the number of connected components of the graph.
minGroupCount <= maxGroupCountminGroupCount <= graph.N()maxGroupCount > 0O(graph.E()) * O(edgeBetweenness)
In practice, it is faster because edge betweenness is computed for subgraphs during the process and this
algorithm terminates after maxGroupCount groups have been determined.
graph - the input graphclusterIDs - the NodeMap that will be filled during the execution and returns an integer value
(cluster ID) for each nodedirected - true if the graph should be considered as directed, false otherwiseminGroupCount - the minimum number of groups that will be returnedmaxGroupCount - the maximum number of groups that will be returnededgeCosts - the DataProvider that holds a positive Double cost or null if
the edges of the graph are considered to be of equal cost
java.lang.IllegalArgumentException - if minGroupCount > maxGroupCount or
minGroupCount > graph.N() or
maxGroupCount <= 0
public static int edgeBetweennessClustering(Graph graph,
NodeMap clusterIDs,
double qualityTimeRatio,
int minGroupCount,
int maxGroupCount,
boolean refine)
In each iteration the edge with the highest betweenness centrality is removed from the graph. The method stops, if there are no more edges to remove or if the requested maximum number of groups is found. The clustering with the best quality reached during the process is returned.
The algorithm includes several heuristic speed-up techniques available through the quality/time ratio. For the
highest quality setting, it is used almost unmodified. The fast betweenness approximation of Brandes and Pich
(Centrality Estimation in Large Networks) is employed for values around 0.5. Typically, this
results in a tiny decrease in quality but a large speed-up and is the recommended setting. To achieve the lowest
running time, a local betweenness calculation is used
(Gregory: Local Betweenness for Finding Communities in Networks).
The method requires the maximum number of groups that will be returned. The smaller this value is, the
faster the overall computation time.
The upper bound on the number of groups is graph.N(). Also, the number of returned groups is
never smaller than the number of connected components of the graph.
minGroupCount <= maxGroupCountminGroupCount <= graph.N()maxGroupCount > 0O(graph.E()) * O(edgeBetweenness)
In practice, it is faster because edge betweenness is computed for subgraphs during the process and this
algorithm terminates after maxGroupCount groups have been determined.
graph - the input graphclusterIDs - the NodeMap that will be filled during the execution and returns an integer value
(cluster ID) for each nodequalityTimeRatio - a value between 0.0 (low quality, fast) and 1.0 (high quality, slow);
the recommended value is 0.5minGroupCount - the minimum number of groups that will be returnedmaxGroupCount - the maximum number of groups that will be returnedrefine - true if the algorithm refines the current grouping, false if the
algorithm discards the current grouping
java.lang.IllegalArgumentException - if minGroupCount > maxGroupCount or
minGroupCount > graph.N() or
maxGroupCount <= 0
public static int biconnectedComponentGrouping(Graph graph,
NodeMap groupIDs)
Nodes will be grouped such that the nodes within each group are biconnected. Nodes that belong to multiple biconnected components will be assigned to exactly one of these components.
groupID for such a node will be null.O(graph.E() + graph.N())graph - the input graphgroupIDs - the NodeMap that will be filled during the execution and returns an integer value (cluster ID)
for each node
public static int labelPropagation(Graph graph,
NodeMap finalLabel)
This algorithm iteratively assigns a label to each node. The label of a node is set to the most frequent
label among its neighbors. If there are multiple candidates the algorithm randomly chooses one of them.
After applying the algorithm, all nodes with the same label (i.e. nodes for which NodeMap
finalLabel returns the same integer value) belong to the same community.
At the start of the algorithm, each node has a unique label. Use method
labelPropagation(Graph, NodeMap, DataProvider) or
labelPropagation(Graph, NodeMap, DataProvider, DataProvider, DataProvider, DataProvider)
to specify custom initial labels.
labelPropagation(Graph, NodeMap, DataProvider, DataProvider, DataProvider, DataProvider) that
allows to specify edge weights. Setting the weight of self-loops to 0.0 results in ignoring them.graph - the undirected input graphfinalLabel - the NodeMap that returns the integer labels of each node after applying the algorithm
where nodes without a label receive the value -1
public static int labelPropagation(Graph graph,
NodeMap finalLabel,
DataProvider initialLabel)
This algorithm iteratively assigns a label to each node. The label of a node is set to the most frequent
label among its neighbors. If there are multiple candidates the algorithm randomly chooses one of them.
After applying the algorithm, all nodes with the same label (i.e. nodes for which NodeMap
finalLabel returns the same integer value) belong to the same community.
At the start of the algorithm, each node is associated with the label returned by method DataProvider.getInt(Object)
of initialLabel. Negative values are ignored, i.e., at the start there may be nodes without label.
This implies that there may be results, where some of the nodes are not associated with labels, too.
For such nodes method NodeMap.getInt(Object) of finalLabel returns -1.
labelPropagation(Graph, NodeMap, DataProvider, DataProvider, DataProvider, DataProvider) that allows
to specify edge weights. Setting the weight of self-loops to 0.0 results in ignoring them.graph - the undirected input graphfinalLabel - the NodeMap that returns the integer labels of each node after applying the algorithm
where nodes without a label receive the value -1initialLabel - DataProvider that stores the initial integer labels of each node (negative values
indicate that a node is not associated with an initial label)
public static int labelPropagation(Graph graph,
NodeMap finalLabel,
DataProvider initialLabel,
DataProvider nodeWeight,
DataProvider edgeWeight,
DataProvider edgeDirectedness)
This algorithm iteratively assigns a label to each node. The label of a node is set to the most frequent
label among its neighbors. If there are multiple candidates the algorithm randomly chooses one of them.
After applying the algorithm, all nodes with the same label (i.e. nodes for which NodeMap
finalLabel returns the same integer value) belong to the same community.
For a given node and neighbor, the weight of the neighbors label is
nodeWeight(neighbor) * edgeWeight((node, neighbor)). In addition, the overall weight of a
specific label is the sum of such weights for all neighbors with matching label.
At the start of the algorithm, each node is associated with the label returned by method DataProvider.getInt(Object)
of initialLabel. Negative values are ignored, i.e., at the start there may be nodes without label.
This implies that there may be results, where some of the nodes are not associated with labels, too.
For such nodes method NodeMap.getInt(Object) of finalLabel returns -1.
If DataProvider edgeDirectedness is specified, the algorithm only considers a subset of the
neighbors when determining the label of a node. More precisely, it only considers the neighbors connected
by edges to a node n for which
0.0 and the edge is incoming (i.e. n is the edge's target),
0.0 and the edge is outgoing (i.e. n is the edge's source),
0.0 (i.e. the edge is considered to be undirected).
0.0 results in ignoring them.graph - the undirected input graphfinalLabel - the NodeMap that returns the integer labels of each node after applying the algorithm
where nodes without a label receive the value -1initialLabel - DataProvider that stores the initial integer labels of each node (negative values
indicate that a node is not associated with an initial label)nodeWeight - DataProvider that stores the weights of the nodesedgeWeight - DataProvider that stores the weights of the edgesedgeDirectedness - DataProvider that stores the directedness of the edges
public static int louvainModularity(Graph graph,
NodeMap communityIndex)
The algorithm starts by assigning each node to its own community. Then, iteratively tries to construct communities by moving nodes from their current community to another until the modularity is locally optimized. At the next step, the small communities found are merged to a single node and the algorithm starts from the beginning until the modularity of the graph cannot be further improved.
The community index of a node corresponds to the index of the associated community. If there are nodes that are
not associated with a community, their index is -1.
1.0. Use method
louvainModularity(Graph, NodeMap, DataProvider) to specify custom edge weights.graph - the input graphcommunityIndex - NodeMap that stores, for each node, the integer community index of the node
public static int louvainModularity(Graph graph,
NodeMap communityIndex,
DataProvider edgeWeight)
The algorithm starts by assigning each node to its own community. Then, iteratively tries to construct communities by moving nodes from their current community to another until the modularity is locally optimized. At the next step, the small communities found are merged to a single node and the algorithm starts from the beginning until the modularity of the graph cannot be further improved.
The community index of a node corresponds to the index of the associated community. If there are nodes that are
not associated with a community, their index is -1.
If no DataProvider for the edge weights is given, the algorithm will assume that all edges have edge weights
equal to 1.
graph - the input graphcommunityIndex - NodeMap that stores, for each node, the integer community index of the nodeedgeWeight - DataProvider that stores the weights of the edges
public static double getModularity(Graph graph,
DataProvider communityIndex)
The modularity value estimates the quality of the partition of the nodes into the given communities and is a value
between [-1/2,1]. High modularity values denote that nodes with dense connections are placed in the same
community, while nodes with sparse connections are assigned to different communities.
To compute the modularity, a DataProvider communityIndex is needed that returns an
integer value representing the community index for each node.
1.0.
Use method getModularity(Graph, DataProvider, DataProvider) to specify custom edge weights.graph - the input graphcommunityIndex - the community index for each node (nodes with same index are associated with the same community)
java.lang.IllegalArgumentException - if no DataProvider communityIndex for the community indices is specifiedlouvainModularity(Graph, NodeMap),
labelPropagation(Graph, NodeMap)
public static double getModularity(Graph graph,
DataProvider communityIndex,
DataProvider edgeWeight)
The modularity value estimates the quality of the partition of the nodes into the given communities and is a value
between [-1/2,1]. High modularity values denote that nodes with dense connections are placed in the same
community, while nodes with sparse connections are assigned to different communities.
To compute the modularity, a DataProvider communityIndex is needed that returns an
integer value representing the community index for each node.
If no DataProvider for the edge weights is given, the algorithm will do the calculation assuming that all
edges have edge weight equals to 1.0. Note that in the case where, all edges have edge weights
equal to 0.0, the modularity will be 0.0, too.
graph - the input graphcommunityIndex - the community index for each node (nodes with same index are associated with the same community)edgeWeight - the weight of the edges
java.lang.IllegalArgumentException - if no DataProvider communityIndex for the community indices is specifiedlouvainModularity(Graph, NodeMap, DataProvider),
labelPropagation(Graph, NodeMap, DataProvider, DataProvider, DataProvider, DataProvider)
public static Groups.Dendrogram hierarchicalClustering(Graph graph,
Groups.Distances distances,
byte linkage)
The clustering is performed using the agglomerative strategy i.e., a bottom-up approach according to which at the beginning each node belongs to its own cluster. At each step pairs of clusters are merged while moving up to the hierarchy. The dissimilarity between clusters is determined based on the given linkage and the given node distances metric. The algorithm continues until all nodes belong to the same cluster.
The result is returned as a Groups.Dendrogram object which represents the result of the clustering
algorithm as a binary tree structure. It can easily be traversed by starting from the
root node and moving on to nodes of the next level via method
Groups.Dendrogram.getChildren(Node).
Groups.Distances object returns a negative distance value for a pair of nodes, zero will be used instead.O(graph.N() ^ 3)graph - the input graphdistances - a given Groups.Distances object that determines the distance between any two nodeslinkage - one of the predefined linkage values
Groups.Dendrogram which represents the result of the clustering as a binary tree or
null if the given graph is empty
java.lang.IllegalArgumentException - if an unknown linkage is given
public static int hierarchicalClustering(Graph graph,
NodeMap clusterIDs,
Groups.Distances distances,
byte linkage,
double cutOff)
The clustering is performed using the agglomerative strategy i.e., a bottom-up approach according to which at the beginning each node belongs to its own cluster. At each step pairs of clusters are merged while moving up to the hierarchy. The dissimilarity between clusters is determined based on the given linkage and the given node distances. The algorithm continues until all nodes belong to the same cluster.
The result will be given based on the given cut-off value that is used for cutting the hierarchical tree at a point such that the dissimilarity values of the nodes that remain at the dendrogram are less than this value.
O(graph.N() ^ 3)graph - the input graphclusterIDs - the NodeMap that will be filled during the execution and returns an integer value (cluster ID)
for each nodedistances - a given Groups.Distances object that determines the distance between any two nodeslinkage - one of the predefined linkage valuescutOff - the cut-off value that determines where to cut the hierarchic tree into clusters
java.lang.IllegalArgumentException - if an unknown linkage is used
public static int hierarchicalClustering(Graph graph,
int maxCluster,
NodeMap clusterIDs,
Groups.Distances distances,
byte linkage)
The clustering is performed using the agglomerative strategy i.e., a bottom-up approach according to which at the beginning each node belongs to its own cluster. At each step pairs of clusters are merged while moving up to the hierarchy. The dissimilarity between clusters is determined based on the given linkage and the given node distances. The algorithm continues until all nodes belong to the same cluster.
The result will be given based on the given maximum number of clusters value that is used for cutting the hierarchical tree at a point such that the number of remaining clusters equals to this value.
The maximum number of clusters needs to be greater than zero and less than the number of the nodes of the graph.
Groups.Distances object returns a negative distance value for a pair of nodes, zero will be used instead.O(graph.N() ^ 3)graph - the input graphmaxCluster - the maximum number of clusters that determines where to cut the hierarchic tree into clustersclusterIDs - the NodeMap that will be filled during the execution and returns an integer value (cluster ID)
for each nodedistances - a given Groups.Distances object that determines the distance between any two graph nodeslinkage - one of the predefined linkage values
java.lang.IllegalArgumentException - if an unknown linkage is given or if the maximum number of clusters is less
than or equal to zero or greater than the number of nodes of the graph
public static int kMeansClustering(Graph graph,
NodeMap clusterIDs,
DataProvider nodePositions,
byte distanceMetric,
int k,
int iterations,
YPoint[] centroids)
The nodes of the graph will be partitioned in k clusters based on their positions such that their
distance from the cluster's mean (centroid) is minimized.
The distance can be defined using diverse metrics as euclidean distance, euclidean-squared distance, manhattan distance or chebychev distance.
k will
be set equal to 2.k or if no centroids are given, random
initial centroids will be assigned for all clusters.O(graph.N() * k * d * I) where k is the number of clusters,
I the maximum number of iterations and d the dimension of the pointsgraph - the input graphclusterIDs - the NodeMap that will be filled during the execution and returns an integer value (cluster ID)
for each nodenodePositions - the DataProvider that holds a point representing the current position of each
node in the graphdistanceMetric - one of the predefined distance metricsk - the number of clustersiterations - the maximum number of iterations performed by the algorithm for convergencecentroids - the initial centroids
java.lang.IllegalArgumentException - if the given distance metric is not supported
public static int kMeansClustering(Graph graph,
NodeMap clusterIDs,
DataProvider nodePositions,
byte distanceMetric,
int k)
kMeansClustering(Graph, NodeMap, DataProvider, byte, int, int, YPoint[]), but no initial centroids are
given as input.
k will
be set equal to 2.100.O(graph.N() * k * d * I) where k is the number of clusters,
I the maximum number of iterations and d the dimension of the pointsgraph - the input graphclusterIDs - the NodeMap that will be filled during the execution and returns an integer value (cluster ID)
for each nodenodePositions - the DataProvider that holds a point representing the current position of each
node in the graphdistanceMetric - one of the predefined distance metricsk - the number of clusters
java.lang.IllegalArgumentException - if the given distance metric is not supported
public static double getClusteringCoefficient(Graph graph,
NodeMap coefficientMap,
boolean directed)
The clustering coefficient measures the degree to which the nodes of a network tend to cluster together,
see https://en.wikipedia.org/wiki/Clustering_coefficient.
More precisely, for a node n, the local clustering coefficient is the actual number of edges between the neighbors
of n divided by the maximum possible number of such edges.
Hence, it is always a Double value between 0.0 and 1.0.
graph - the input graphcoefficientMap - a map that returns the clustering coefficient for each nodedirected - whether or not the graph is directed
|
© Copyright 2000-2025, yWorks GmbH. All rights reserved. |
||||||||
| PREV CLASS NEXT CLASS | FRAMES NO FRAMES | ||||||||
| SUMMARY: NESTED | FIELD | CONSTR | METHOD | DETAIL: FIELD | CONSTR | METHOD | ||||||||