 y.algo.Centrality
y.algo.Centrality
|
Search this API | ||||||||
| PREV CLASS NEXT CLASS | FRAMES NO FRAMES | ||||||||
| SUMMARY: NESTED | FIELD | CONSTR | METHOD | DETAIL: FIELD | CONSTR | METHOD | ||||||||
java.lang.Objecty.algo.Centrality
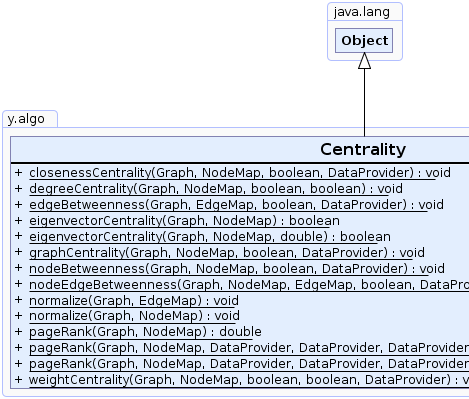
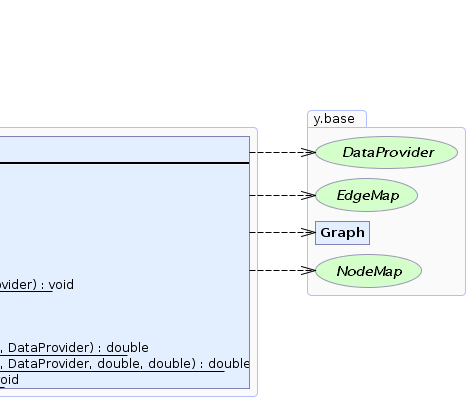
public class Centrality
This class provides methods to determine various centrality indices of nodes or edges of a graph.
Centrality indices serve to quantify an intuitive feeling that in most networks some nodes or edges are "more central" than others. The provided methods assign a double value to each node or edge of a graph that represents its centrality. The higher an assigned value, the more central the element is considered by the algorithm.
Also, this class provides convenience methods that normalize the returned centrality values such that they lie within
the interval [0,1].
 |
 |
| Method Summary | |
|---|---|
static void |
closenessCentrality(Graph graph,
NodeMap closeness,
boolean directed,
DataProvider edgeCosts)
Computes the closeness centrality for the nodes of a graph. |
static void |
degreeCentrality(Graph graph,
NodeMap centrality,
boolean considerInEdges,
boolean considerOutEdges)
Computes the degree centrality for the nodes of a given graph. |
static void |
edgeBetweenness(Graph graph,
EdgeMap centrality,
boolean directed,
DataProvider edgeCosts)
Computes betweenness centrality for each edge of a given graph. |
static boolean |
eigenvectorCentrality(Graph graph,
NodeMap centralityMap)
Computes an eigenvector centrality for each node of a given undirected, unweighted graph. |
static boolean |
eigenvectorCentrality(Graph graph,
NodeMap centralityMap,
double precision)
Computes an eigenvector centrality for each node of a given undirected graph. |
static void |
graphCentrality(Graph graph,
NodeMap centrality,
boolean directed,
DataProvider edgeCosts)
Computes the graph centrality for the nodes of a graph. |
static void |
nodeBetweenness(Graph graph,
NodeMap centrality,
boolean directed,
DataProvider edgeCosts)
Computes betweenness centrality for each node of a given graph. |
static void |
nodeEdgeBetweenness(Graph graph,
NodeMap nodeCentrality,
EdgeMap edgeCentrality,
boolean directed,
DataProvider edgeCosts)
Computes betweenness centrality for each node and edge of a given graph. |
static void |
normalize(Graph graph,
EdgeMap map)
Normalizes the double values of a given EdgeMap by dividing each of them by the
maximum of all values (maximum norm). |
static void |
normalize(Graph graph,
NodeMap map)
Normalizes the double values of a given NodeMap by dividing each of them by the
maximum of all values (maximum norm). |
static double |
pageRank(Graph graph,
NodeMap pageRank)
Computes page rank values for all nodes based on their attached edges. |
static double |
pageRank(Graph graph,
NodeMap pageRank,
DataProvider initialPageRank,
DataProvider nodeWeight,
DataProvider edgeWeight,
DataProvider edgeDirectedness)
Computes page rank values for all nodes based on their attached edges. |
static double |
pageRank(Graph graph,
NodeMap pageRank,
DataProvider initialPageRank,
DataProvider nodeWeight,
DataProvider edgeWeight,
DataProvider edgeDirectedness,
double dampingFactor,
double precision)
Computes page rank values for all nodes based on their attached edges. |
static void |
weightCentrality(Graph graph,
NodeMap centrality,
boolean considerInEdges,
boolean considerOutEdges,
DataProvider edgeWeights)
Computes the weight centrality for the nodes of a graph. |
| Methods inherited from class java.lang.Object |
|---|
clone, equals, finalize, getClass, hashCode, notify, notifyAll, toString, wait, wait, wait |
| Method Detail |
|---|
public static void nodeBetweenness(Graph graph,
NodeMap centrality,
boolean directed,
DataProvider edgeCosts)
Betweenness centrality is a measure for how often a node lies on a shortest path between each pair of nodes in the graph. Removing a central edge will cause many shortest paths to change.
O(graph.N() * graph.E()) for unweighted graphsO(graph.N() * (graph.E() + graph.N()) * log(graph.N()) for weighted graphsgraph - the input graphcentrality - the NodeMap that will be filled during the execution and returns a double value (centrality)
for each nodedirected - true if the graph should be considered as directed, false otherwiseedgeCosts - the DataProvider that returns a positive double value (cost) or null if the edges
are of equal cost; for invalid input values the algorithm uses cost 1.0
public static void edgeBetweenness(Graph graph,
EdgeMap centrality,
boolean directed,
DataProvider edgeCosts)
Betweenness centrality is a measure for how often an edge lies on a shortest path between each pair of nodes in the graph. Removing a central edge will cause many shortest paths to change.
O(graph.N() * graph.E()) for unweighted graphsO(graph.N() * (graph.E() + graph.N()) * log(graph.N()) for weighted graphsgraph - the input graphcentrality - the EdgeMap that will be filled during the execution and returns a double value (centrality) for
each edgedirected - true if the graph should be considered as directed, false otherwiseedgeCosts - the DataProvider that returns a positive double value (cost) or null if the edges
are of equal cost; for invalid input values the algorithm uses cost 1.0
public static void nodeEdgeBetweenness(Graph graph,
NodeMap nodeCentrality,
EdgeMap edgeCentrality,
boolean directed,
DataProvider edgeCosts)
Betweenness centrality is a measure for how often a node/edge lies on a shortest path between each pair of nodes in the graph. Removing a central node/edge will cause many shortest paths to change.
O(graph.N() * graph.E()) for unweighted graphsO(graph.N() * (graph.E() + graph.N()) * log(graph.N()) for weighted graphsgraph - the input graphnodeCentrality - the NodeMap that will be filled during the execution and returns a double value
(centrality) for each nodeedgeCentrality - the EdgeMap that will be filled during the execution and returns a double value
(centrality) for each edgedirected - true if the graph should be considered as directed, false otherwiseedgeCosts - the DataProvider that returns a positive double value (cost) or null if the edges
are of equal cost; for invalid input values the algorithm uses cost 1.0
public static void closenessCentrality(Graph graph,
NodeMap closeness,
boolean directed,
DataProvider edgeCosts)
Closeness centrality is defined as the reciprocal of the sum of shortest path distances of a node to all other
nodes in the graph. Therefore, a node with high closeness centrality has short distances to all other nodes
of a graph. If the sum of the shortest path distances is 0, the closeness of a node is set to
Double.POSITIVE_INFINITY.
connected.O(graph.N()^2 + graph.N() * graph.E()) for unweighted graphsO((graph.N() * graph.E()) + graph.N()^2 * log(graph.N())) for weighted graphsgraph - the input graphcloseness - the NodeMap that will be filled during the execution and returns a double value
(centrality) for each nodedirected - true if the graph should be considered as directed, false otherwiseedgeCosts - the DataProvider that returns a positive double value (cost) or null if the edges
are of equal cost
public static void graphCentrality(Graph graph,
NodeMap centrality,
boolean directed,
DataProvider edgeCosts)
Graph centrality is defined as the reciprocal of the maximum of all shortest path distances from a node to all other nodes in the graph. Nodes with high graph centrality have short distances to all other nodes in the graph.
NodeMap must be given as input that returns for each Node a zero value
that represents the initial closeness centrality.O(graph.N()^2 + graph.N() * graph.E()) for unweighted graphsO((graph.N() * graph.E()) + graph.N()^2 * log(graph.N())) for weighted graphsgraph - the input graphcentrality - the NodeMap that will be filled during the execution and returns a double value
(centrality) for each nodedirected - true if the graph should be considered as directed, false otherwiseedgeCosts - the DataProvider that returns a positive double value (cost) or null if the edges
are of equal cost
public static void degreeCentrality(Graph graph,
NodeMap centrality,
boolean considerInEdges,
boolean considerOutEdges)
Degree centrality is the number of the incoming, outgoing or overall edges incident to a node (measures incoming, outgoing and overall degree).
considerInEdges and considerOutEdges must be
true.O(graph.N())graph - the input graphcentrality - the NodeMap that will be filled during the execution and returns a double value
(centrality) for each nodeconsiderInEdges - true if the incoming edges should be considered, false otherwiseconsiderOutEdges - true if the outgoing edges should be considered, false otherwise
public static void weightCentrality(Graph graph,
NodeMap centrality,
boolean considerInEdges,
boolean considerOutEdges,
DataProvider edgeWeights)
Weight centrality measures the weight associated with incoming, outgoing, or all edges of a node.
edgeWeights is null, then
degreeCentrality is invoked instead.O(graph.E())graph - the input graphcentrality - the NodeMap that will be filled during the execution and returns a double value
(centrality) for each nodeconsiderInEdges - true if the incoming edges should be considered, false otherwiseconsiderOutEdges - true if the outgoing edges should be considered, false otherwiseedgeWeights - the DataProvider that returns a positive double value (weight) or null if the edges
are considered to have uniform weight of 1.0
public static void normalize(Graph graph,
NodeMap map)
double values of a given NodeMap by dividing each of them by the
maximum of all values (maximum norm).
Double.POSITIVE_INFINITY, all values other
than Double.POSITIVE_INFINITY are set to 0.n: map.getDouble(n) >= 0graph - the input graphmap - the NodeMap that will be filled during the execution and returns a double value
from [0,1] interval
public static void normalize(Graph graph,
EdgeMap map)
double values of a given EdgeMap by dividing each of them by the
maximum of all values (maximum norm).
Double.POSITIVE_INFINITY, all values other
than Double.POSITIVE_INFINITY are set to 0.e: map.getDouble(e) >= 0graph - the input graphmap - the EdgeMap that will be filled during the execution and returns a double value
from [0,1] interval
public static boolean eigenvectorCentrality(Graph graph,
NodeMap centralityMap)
The centrality values are scaled so that the largest centrality value is 1.0.
PageRank algorithm is also based on eigenvector centrality but allows for more configuration options.graph - the undirected input graphcentralityMap - a node map that can store values of type double. It is used for returning the
resulting centrality values (in the interval [0,1]) for each node
centralityMap contains valid entries
public static boolean eigenvectorCentrality(Graph graph,
NodeMap centralityMap,
double precision)
The centrality values are scaled so that the largest centrality value is 1.0.
PageRank algorithm.graph - the undirected input graphcentralityMap - a node map that can store values of type double. It is used for returning the
resulting centrality values (in the interval [0,1]) for each nodeprecision - specifies the precision used during the calculation of the power iteration method
(i.e. the maximum possible difference to consider two values as equal)
centralityMap contains valid entries
public static double pageRank(Graph graph,
NodeMap pageRank)
The original algorithm was proposed by Sergey Brin and Lawrence Page as PageRank algorithm and refined
later in several papers:
S. Brin and L. Page. The anatomy of a large-scale hypertextual Web search engine.
Computer Networks and ISDN Systems, 30(1-7):107-117
1998.
O(graph.N() + graph.E())graph - the graph to calculate the ranks forpageRank - the NodeMap that will be filled during the execution with the rank value for each node
public static double pageRank(Graph graph,
NodeMap pageRank,
DataProvider initialPageRank,
DataProvider nodeWeight,
DataProvider edgeWeight,
DataProvider edgeDirectedness)
The original algorithm was proposed by Sergey Brin and Lawrence Page as PageRank algorithm and refined
later in several papers:
S. Brin and L. Page. The anatomy of a large-scale hypertextual Web search engine.
Computer Networks and ISDN Systems, 30(1-7):107-117
1998.
The algorithm starts by initializing the rank value for each node (see DataProvider initialPageRank).
After that it uses multiple iterations to propagate the rank of a node to its successors.
The base factor for each successor is 1. These base factors are multiplied by edge weights
(see data provider edgeWeight and node weights
(see data provider nodeWeight if specified.
The final node factors are scaled afterwards to sum up to 1 so the overall rank stays
the same after each iteration. The old page rank of the node multiplied with these scaled factors are then distributed
to the successors.
When a node can't distribute its rank, e.g. because it has no successors, its rank is
distributed between all nodes scaled by their weight (see data provider nodeWeight).
After each iteration, the relative change of each node's page rank ((newRank - oldRank)/oldRank) is calculated. If all page rank changes are below some threshold or if the maximal iterations are reached, the algorithm stops.
The edgeDirectedness DataProvider defines the direction of the edges. This in consequence
defines the propagation direction of ranks. Edges with a positive directedness value propagate the rank from the
source to the target node - this is also the default if the given data provider is null.
Edges which have a directedness of 0 are considered to be undirected so that propagation happens
from source to target and from target to source. Edges with a negative directedness are considered
to have the reverse direction such that they propagate ranks only from target to source.
DataProvider initialPageRank is null, the algorithm assumes that all
nodes have an initial rank of 1.0.O(graph.N() + graph.E())graph - the graph to calculate the ranks forpageRank - the NodeMap that will be filled during the execution with the rank value for each nodeinitialPageRank - The DataProvider that specifies the initial page rank for each nodenodeWeight - the DataProvider that specifies the weight for each nodeedgeWeight - the DataProvider that specifies the weight for each edgeedgeDirectedness - DataProvider that stores the directedness of the edges
public static double pageRank(Graph graph,
NodeMap pageRank,
DataProvider initialPageRank,
DataProvider nodeWeight,
DataProvider edgeWeight,
DataProvider edgeDirectedness,
double dampingFactor,
double precision)
The original algorithm was proposed by Sergey Brin and Lawrence Page as PageRank algorithm and refined
later in several papers:
S. Brin and L. Page. The anatomy of a large-scale hypertextual Web search engine.
Computer Networks and ISDN Systems, 30(1-7):107-117
1998.
The algorithm starts by initializing the rank value for each node (see DataProvider initialPageRank).
After that it uses multiple iterations to propagate the rank of a node to its successors.
The base factor for each successor is 1. These base factors are multiplied by edge weights
(see data provider edgeWeight and node weights
(see data provider nodeWeight if specified.
The final node factors are scaled afterwards to sum up to 1 so the overall rank stays
the same after each iteration. The old page rank of the node multiplied with these scaled factors are then distributed
to the successors.
When a node can't distribute its rank, e.g. because it has no successors, its rank is
distributed between all nodes scaled by their weight (see data provider nodeWeight).
Furthermore a damping factor is used so that a
share of each nodes' rank isn't propagated to the successors but also distributed between all nodes.
After each iteration, the relative change of each node's page rank ((newRank - oldRank)/oldRank) is calculated.
If all page rank changes are below the specified precision or if the
maximal iterations are reached, the algorithm stops.
The edgeDirectedness DataProvider defines the direction of the edges. This in consequence
defines the propagation direction of ranks. Edges with a positive directedness value propagate the rank from the
source to the target node - this is also the default if the given data provider is null.
Edges which have a directedness of 0 are considered to be undirected so that propagation happens
from source to target and from target to source. Edges with a negative directedness are considered
to have the reverse direction such that they propagate ranks only from target to source.
0 and 1 where 0 means all
the rank of node is distributed between all nodes based on their node weight and 1 means all the
rank of a node is only distributed between its successors.DataProvider initialPageRank is null, the algorithm assumes that all
nodes have an initial rank of 1.0.O(graph.N() + graph.E())graph - the graph to calculate the ranks forpageRank - the NodeMap that will be filled during the execution with the rank value for each nodeinitialPageRank - The DataProvider that specifies the initial page rank for each nodenodeWeight - the DataProvider that specifies the weight for each nodeedgeWeight - the DataProvider that specifies the weight for each edgeedgeDirectedness - DataProvider that stores the directedness of the edgesdampingFactor - a value between 0 and 1 denoting the factor of a node's rank that
shall be distributed to its successors (a good default value is 0.85)precision - the non-negative precision of the calculated page ranks (a good default value is 0.001)
|
© Copyright 2000-2025, yWorks GmbH. All rights reserved. |
||||||||
| PREV CLASS NEXT CLASS | FRAMES NO FRAMES | ||||||||
| SUMMARY: NESTED | FIELD | CONSTR | METHOD | DETAIL: FIELD | CONSTR | METHOD | ||||||||